CellRanger
Generated by: George Edward Chlipala
Report date: January 11, 2021
Overview
When you publish manuscripts based on data generated at our facility, we would greatly appreciate an acknowledgement of our efforts. Please cite our facility as follows (for example):
Basic processing of the raw data were performed by the University of Illinois at Chicago Research Informatics Core (UICRIC).
We adhere to a general policy for acknowledgements and authorship as established by the Association for Biomolecular Resource Facilities (ABRF) , and we support the following statement from the ABRF.
The existence of core facilities depends in part on proper acknowledgment in publications. This is an important metric of the value of most core facilities. Proper acknowledgment of core facilities enables them to obtain financial and other support so that they may continue to provide their essential services in the best ways possible. It also helps core personnel to advance in their careers, adding to the overall health of the core facility.
Please contact us for assistance in drafting manuscripts.
Output Files
| File | Description | Type |
|---|---|---|
| sc5p_v2_mm_balbc_T_1k_5gex_matrix/features.tsv.gz | Gene features for sc5p_v2_mm_balbc_T_1k_5gex | result |
| sc5p_v2_mm_balbc_T_1k_5gex_matrix/barcodes.tsv.gz | Feature list (cell barcodes) for sc5p_v2_mm_balbc_T_1k_5gex | result |
| sc5p_v2_mm_balbc_T_1k_5gex_matrix/matrix.mtx.gz | Gene counts in sparse matrix format for sc5p_v2_mm_balbc_T_1k_5gex | result |
| sc5p_v2_mm_balbc_T_1k_5gex.cloupe | Cellranger Cloupe file for sc5p_v2_mm_balbc_T_1k_5gex | result |
| sc5p_v2_mm_balbc_T_1k_t_airr_rearrangement.tsv | Annotated contigs and consensus sequences of VDJ rearrangements in the AIRR format for sc5p_v2_mm_balbc_T_1k_t | result |
| sc5p_v2_mm_balbc_T_1k_t_all_contig_annotations.csv | High-level and detailed annotations of each contig for sc5p_v2_mm_balbc_T_1k_t | result |
| sc5p_v2_mm_balbc_T_1k_t_all_contig_fasta.gz | Sequences for assembled VD(J) contigs for sc5p_v2_mm_balbc_T_1k_t | result |
| sc5p_v2_mm_balbc_T_1k_t_consensus_annotations.csv | High-level and detailed annotations of each clonotype consensus sequence for sc5p_v2_mm_balbc_T_1k_t | result |
| sc5p_v2_mm_balbc_T_1k_t_clonotypes.csv | High-level descriptions of each clonotype for sc5p_v2_mm_balbc_T_1k_t | result |
| sc5p_v2_mm_balbc_T_1k_t_filtered_contig_annotations.csv | High-level annotations of each high-confidence, cellular contig for sc5p_v2_mm_balbc_T_1k_t | result |
| sc5p_v2_mm_balbc_T_1k_t.cloupe | Cellranger Vloupe file for sc5p_v2_mm_balbc_T_1k_t | result |
Details
- Method: Demultiplexing and gene expression quantification for 10X with CellRanger
-
Raw reads are mapped to the reference genome and demultiplexed into single cells using CellRanger.
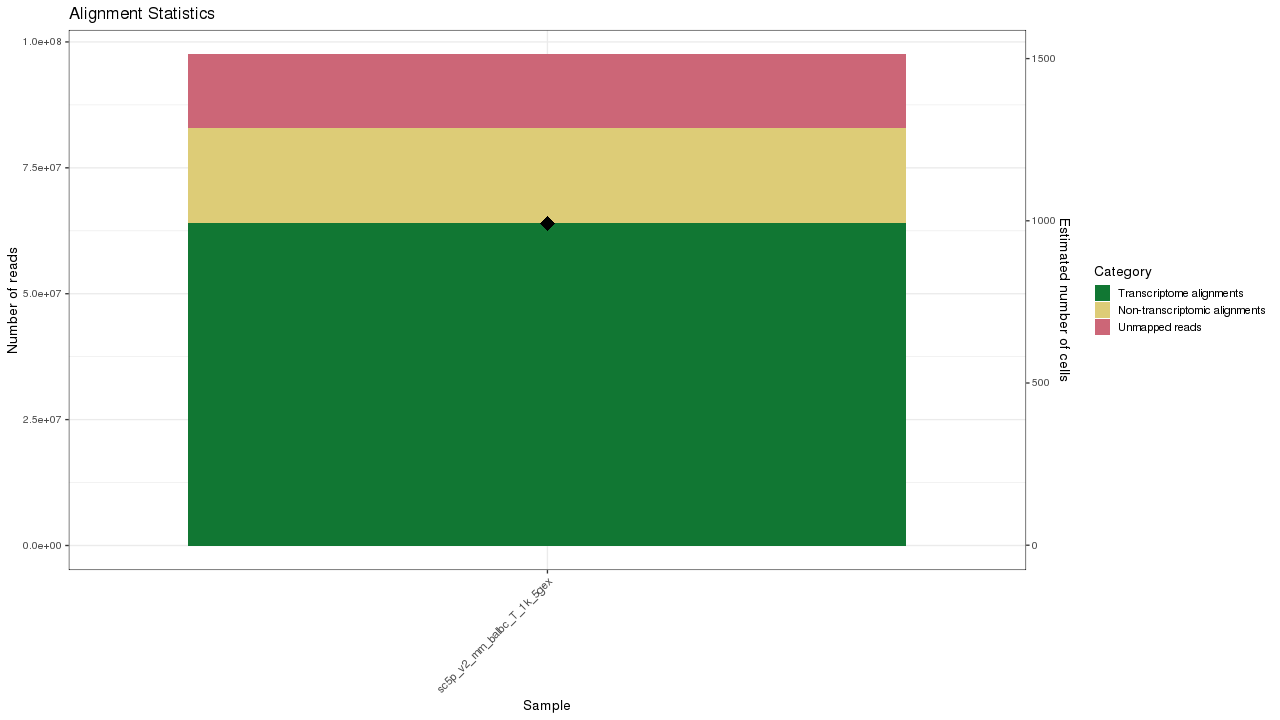
Figure 1. Gene expression (GEX) sample summary
Table 1. Data processing summary statistics of gene expression samples Download table data
| Sample | Estimated Number of Cells | Median UMI Counts per Cell | Median Genes per Cell | Number of Reads | Sequencing Saturation | Valid Barcodes | Reads Mapped Confidently to Genome | Reads Mapped Confidently to Transcriptome |
|---|---|---|---|---|---|---|---|---|
| sc5p_v2_mm_balbc_T_1k_5gex | 992 | 3976 | 1649 | 97505715 | 91.9% | 94.4% | 85.1% | 65.6% |
Details
- Method: Demultiplexing and VD(J) region quantification for 10X with CellRanger
-
Raw reads are mapped to the reference genome and demultiplexed into single cells using CellRanger.
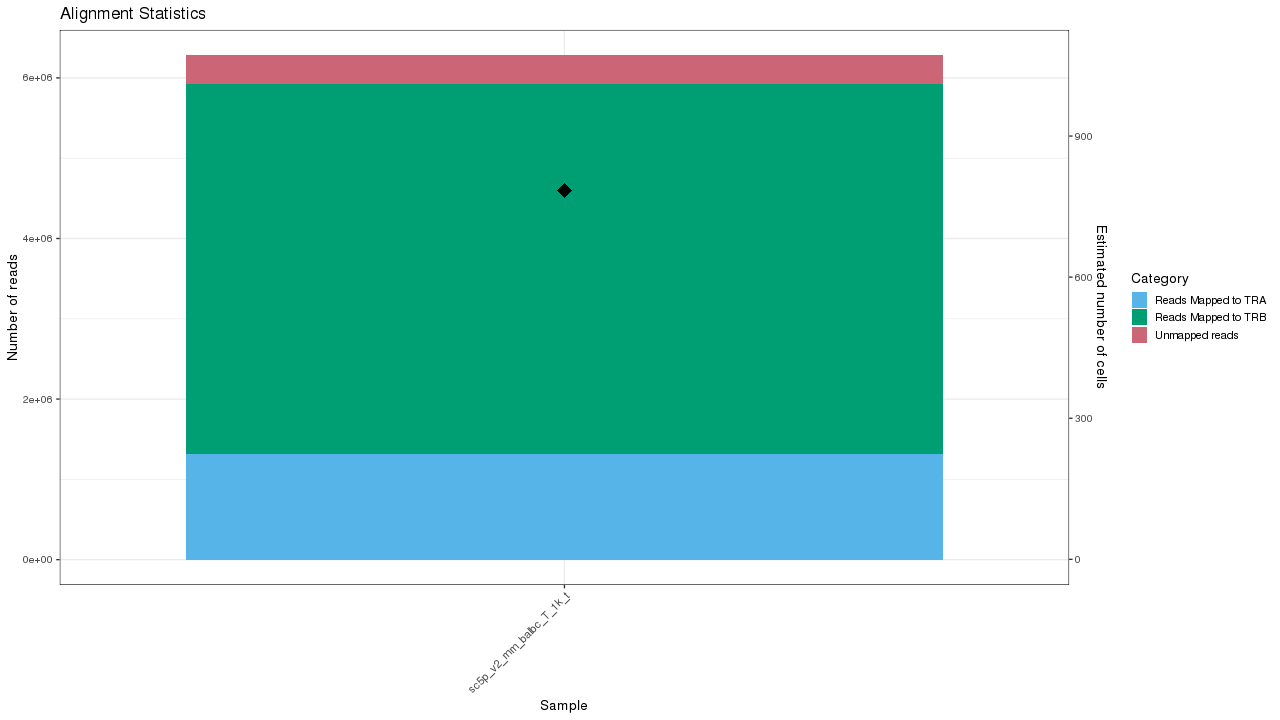
Figure 1. VDJ sample summary
Table 1. Data processing summary statistics of VDJ samples Download table data
| Sample | Estimated Number of Cells | Number of Cells With Productive V-J Spanning Pair | Median TRA UMIs per Cell | Median TRB UMIs per Cell | Number of Read Pairs | Valid Barcodes | Reads Mapped to Any V(D)J Gene | Reads Mapped to TRA | Reads Mapped to TRB |
|---|---|---|---|---|---|---|---|---|---|
| sc5p_v2_mm_balbc_T_1k_t | 785 | 618 | 5 | 11 | 6289407 | 97.4% | 94.2% | 21.0% | 73.1% |